Back in mid-January three members of the University of Sheffield’s Research Software Engineering Team (me, Mike Croucher and Tania Allard) spent a week at a Computational Mathematics with Jupyter workshop, hosted at Edinburgh’s International Centre for Mathematical Sciences.
This brought together the many members of the consortium working on the OpenDreamKit Horizon 2020 European Research Infrastructure project. The overall aim of the project is broad (to further the open-source computational mathematics ecosystem) so it was unsurprising that the collective experience of the attendees was too. The attendees generally fell into one of the following four camps:

The structure was different from conferences I’d attended previously: for each of the five days we listened and debated presentations in the morning then busied ourselves with code sprints in the afternoons.
Our own Mike Croucher kicked things off by asking Is your research software correct? in which he presented Croucher’s Law:
I can be an idiot and will make mistakes.
with the corollary that
You are no different!
He argued, convincingly, that in our research we therefore need to put in place safeguards to lessen the chance and impact of mistakes, and proposed the following as partial solutions:
Raniere Silva from the Software Sustainability Institute followed on with a complementary talk on how to make computational mathematics software more sustainable. He commented that odd numerical bugs can easily creep in over time (e.g. differing floating point behaviour between Python 2 and 3) but that we can maintain confidence in software using version control, continuous integration, good documentation, tutorials, knowledge bases, instant messaging and by developing communities around the software we value.
Alexander Konovalov then talked about a particular case of making research software more sustainable and portable: he’s been using Docker containers to run GAP. This lead into a discussion on whether Docker is a sensible solution for archiving/reproducing workflows: will it be around in ten years’ time? Those interested in that particular issue might benefit from attending the forthcoming Software Sustainability Institute workshop on Docker Containers for Reproducible Research.
We were then given what was pitched as a ‘general introduction’ to Jupyter but ended up covering much more ground than anticipated, largely due to the speaker being Thomas Kluyver, one of the core IPython developers (who happens to have gained his PhD from the University of Sheffield). Thomas talked about the most significant features (literate programming environments; the power and versatility of using the browser as a REPL; Jupyter’s client-server architecture) but also touched upon various tools and platforms that have built on Jupyter including:
Given the enormity of the Jupyter ecosystem and how quickly it has grown it was great to hear from a core developer which related projects he thinks are the most significant and interesting!
Next up, Vidar Fauske gave this talk on nbdime, a new tool for merging and diffing Jupyter Notebooks. The backstory is that for some time we’ve been recommending Jupyter to those wanting to start using Python or R in their research and we’ve also been telling everyone to use version control but the diffing and merging tools typically used with version control systems don’t work well with Notebooks as they
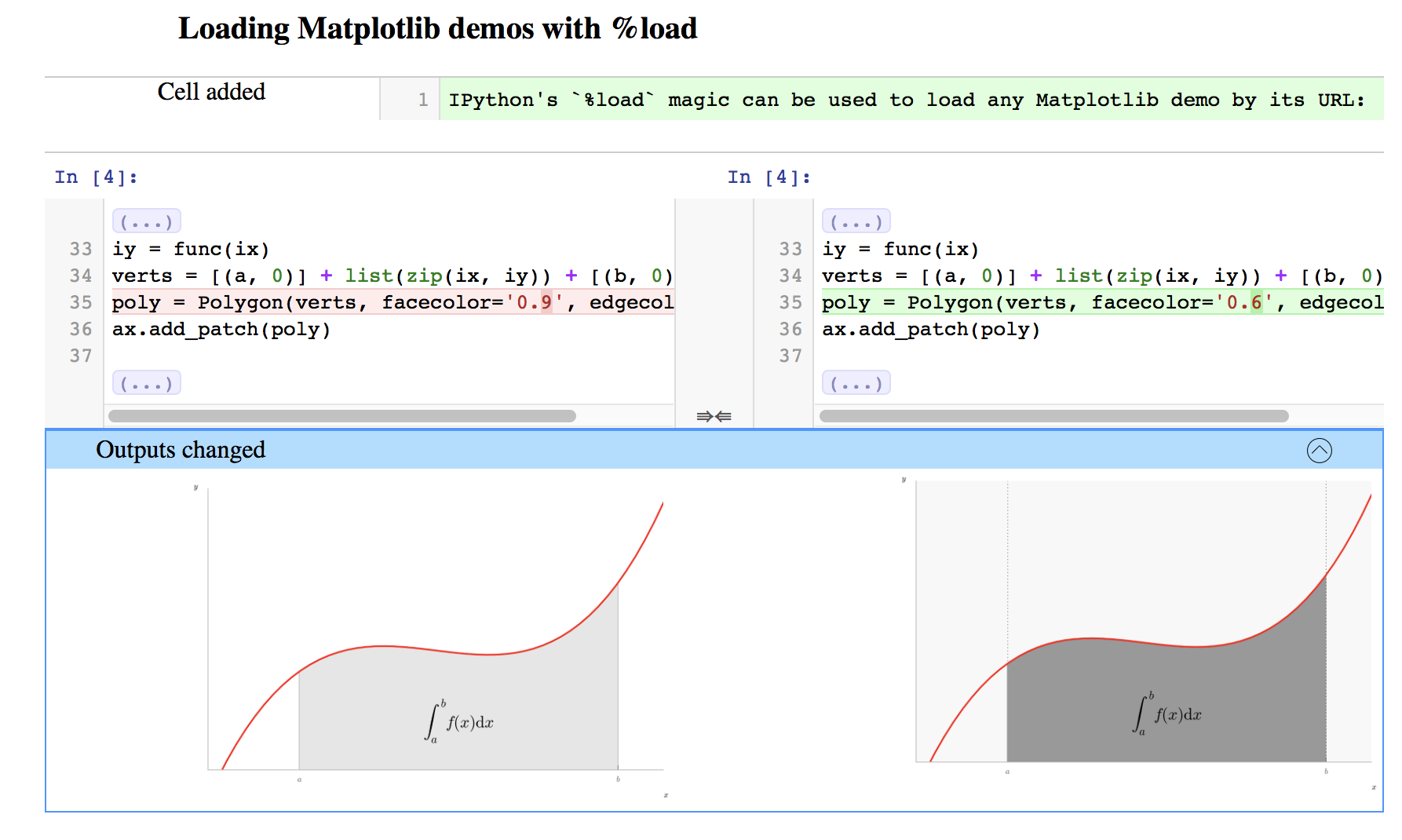
Ultimately visualising the differences between two Notebooks and merging Notebooks in sensible, useful ways really requires that the tools that perform these functions have some understanding of the structure and purpose of Notebooks: nbdime has that awareness:
nbdime provides a core library, plus command-line and browser interfaces for diffing and merging.
Overall, I’m massively excited about nbdime for facilitating much slicker Notebook-based version controlled workflows and hope it sees widespread adoption and promotion by the likes of Software Carpentry.
Hans Fangohr then introduced nbval, a new tool for automating the valdation of Jupyter Notebooks. This could give researchers greater confidence in their workflows: does a demonstrative Notebook still give the same answers if re-run after making changes to the Notebook’s environment (e.g. the package dependencies)?
nbval, a pytest plug-in, works as follows: it creates a copy of a Notebook file, executes the copy in the current Python environment, saves the copy Notebook with its new cell outputs then compares the outputs of the two Notebooks. There are some nice features to control the granularity of testing: flags can be set so certain cells are run but not tested; regexes can be used to ignore oft-changing output strings (e.g. paths, timestamps, memory addresses). Images and LaTeX can’t be handled yet.
Again, I’m exited about this new tool: being able to package both workflow documentation and regression/ acceptance tests as Notebooks is a great idea. Note that at present both nbdime and nbval include mechanisms for comparing Notebooks but are presently separate projects. It will be interesting to see if there’s any convergence in future.
We were treated to two talks on the ipywidgets package, which provides Python and Javascript-backed widgets for interacting with Notebooks e.g. sliders for assessing the impact of model parameters on trends in embedded matplotlib plots.
First, Jeroen Demeyer introduced us to
the high-level interact Python decorator function and interactive
class one can use to control function inputs using a HTML+Javascript
widget. He then went on to explain how one can manually reproduce the
magic of these mechanisms: you instantiate some (typed) input widgets
and output widgets, add them to an on-screen container then associate
each input widget with a callback.
Next, Sylvain Corlay talked about the ipywidgets ecosystem and the future direction of the project. He mentioned several projects that have built on ipywidgets, all of which sound exciting but none of which I’d heard of before this!
The current version of ipywidgets, released since the workshop, includes some interesting developments: much more of the code is now written in Javascript (actually Typescript) rather than Python so widgets state is maintained in JavaScript-land: widgets can therefore now be rendered and manipulated without a Jupyter kernel! See this statically-rendered Notebook on GitHub as an example. Another advantage of migrating the bulk of the code to Javascript is that the widgets should be usable with kernel languages other than Python such as R (once people have written language-specific ipywidgets backends).
Separate to ipywidgets, we were also introduced to SciviJS, a tool currently being developed by Martin Renou at LogiLab for visualising 3D mesh-based geometries in a Juypter Notebook. It uses also uses WebGL / three.js for rendering so is rather performant. I can see some ex-colleagues in civil engineering really liking this. Check out the online demo.
Numbas is a open web-based system for formative and summative maths and science tests. It is being developed by Christian Lawson-Perfect from the University of Newcastle’s Maths and Stats E-Learning unit. It’s very different to teaching environments that use Jupyter (e.g. SageMathCloud) as almost all the code is self-contained HTML+Javascript that is run on the client (for scalability and resilience) and it is for generating closed tests (rather than open mathematical exercises). Looks very attractive and intuitive from the user’s perspective!
Christian also mentioned Up For Grabs, a site of projects wanting help on simpler tasks. He says it’s a good and simple way of getting less experienced developers involved with open-source projects. As a project maintainer you upload some blurb about your project and tell the site which GitHub Issue tag(s) indicate smaller tasks that are ‘up for grabs’.
Hans Fangohr from the University of Southampton reported on using Python and Jupyter to encapsulate multi-stage micro-magnetism modelling workflows: his team have been able to automate the generation of input files and processing of output files for/from old but robust modelling software (OOMMF); Jupyter then further masks away the complexities of running models.
Mark Quinn then talked about the impact that SageMathCloud, an online teaching environment which uses Jupyter, has had on the teaching of physics, astronomy and coding at the University of Sheffield. He’s been working with Mike Croucher to develop SageMathCloud courses for the Physics department with the goal of introducing effective programming tuition early in undergraduate Physics degree programmes. He’s now quite a fan of the used coding environment (Jupyter) and SageMathCloud’s courseware tools (chat facilities and mechanisms for setting and grading assignments) but has now been using it long enough to identify some challenges/issues too (e.g. students getting confused about the order of execution of cells; students opening many notebooks at once (each of which has a resource footprint).
Mark is involved with the Shepherd Group, who research the efficacy of teaching methods and are based in the same Physics department. They’ve recently been studying the impact of using the Jupyter Notebook to undergraduate students who had and hadn’t studied Physics at A-Level. They tested students (at different levels of Bloom’s Taxonomy) before and after teaching and concluded that the Notebooks were suitable for aiding students, regardless of whether they had a Physics background. Hopefully the Software Sustainability Institute can lend their support to pedagogical studies of this nature in future.
I should note that there were also a number of other talks that focussed on the GAP and SageMath computational mathematics software packages: I’ve deliberately not mentioned them here so as not to expose my lack of understanding of group theory and semi-groups and also this post is long enough already! See the full programme for info on things I’ve neglected plus links to the presentations.
After lunch on each of the five days of the workshop we worked on various coding sprints: some wrote a Jupyter kernel for GAP, others worked on SageMath, whilst me and a few others started work on Jupyter Interactions. This sprint was suggested by Mike Croucher: he thought it would be nice to have a curated set of Notebooks that demonstrate how to use different ipywidgets to manipulate various mathematical objects. Several of us wrote Notebooks while Christian Lawson-Perfect quickly put together a very nice gallery-generating front end:
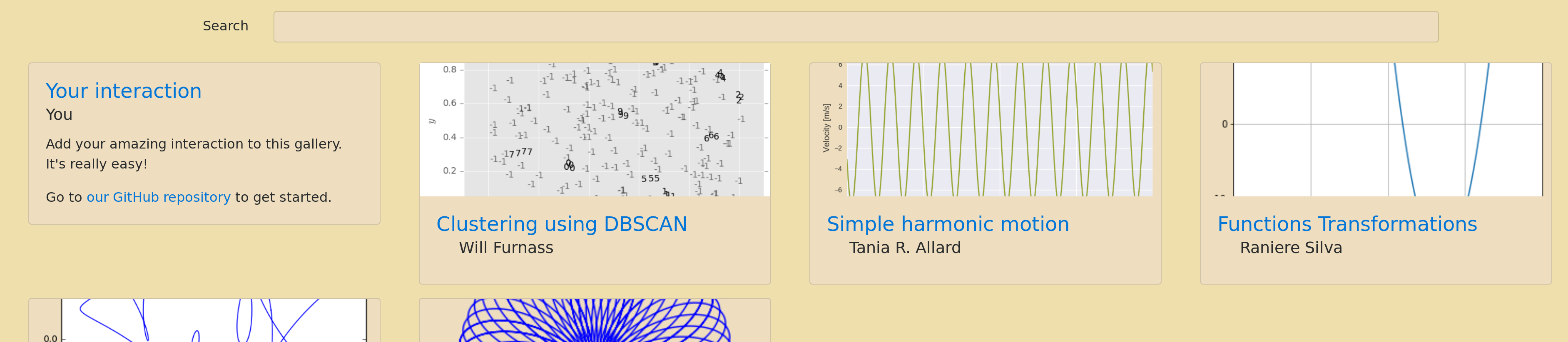
See the end result here. Should you wish to submit your own Jupyter Interactions Notebook then please submit a pull request!
This was the first time I’d been to a conference where the emphasis was very much on sharing ideas and working together: the academic conferences I’d attended prior to this had previously had an air of competition about them. Looking forward to meeting up with the OpenDreamKit gang again!
For queries relating to collaborating with the RSE team on projects: rse@sheffield.ac.uk
Information and access to Bede.
Join our mailing list so as to be notified when we advertise talks and workshops by subscribing to this Google Group.
Queries regarding free research computing support/guidance should be raised via our Code clinic or directed to the University IT helpdesk.