*The Part one of this series, was appeared on the RSE blog.
The University of Sheffield partnered with NVIDIA and Oak Ridge Leadership Computing Facility (OLCF) to hold its first GPU hackathon, where seven applications teams worked alongside mentors with GPU-programming expertise to accelerate their scientific codes using GPUs.
Following the success of the first Sheffield GPU Hackathon, in 2020 we will once again bring together team of application developers some with little or no previous GPU programming experience to get their own codes running on GPUs, or to optimise their codes that currently run on GPUs, using OpenACC or other parallel programming models. Stay tuned for future announcements via http://gpuhackathons.org/!
But first, let’s hear some of the success stories from the teams participated in the 2019 Sheffield GPU Hackathon:
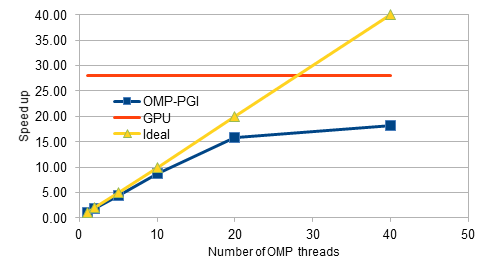
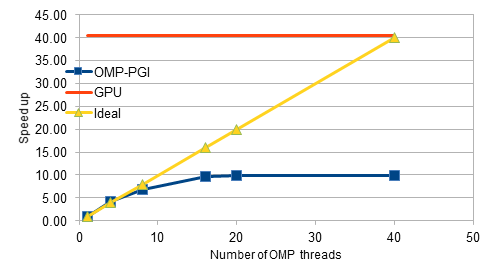
Our project focussed on the parallelisation of an in-house explicit CFD code on GPUs. The code is written in C++ and initially has been parallelised for shared-memory architectures using OpenMP constructs. During the hackathon, the Class objects for the finite elements have been modified by removing the virtual functions, which are not supported yet with OpenACC constructs. The preliminary design changes and restructuring of data allocations has improved the performance of the serial code by about 30%. Later, the parallel code with OpenMP constructs has been tested and profiled for performance, and observed that the scalability flattens out beyond 8 threads. The code is then parallelised for GPUs by adding the OpenACC constructs. After several iterations of profiling, testing and design changes, we achieved an 28X speed-up for the medium-sized mesh consisting of 0.16 million nodes. For the fine mesh with 4 million nodes, we achieved a speed-up of 40X as opposed to the mere 10X obtained with the OpenMP code. Overall, the GPU hackathon has been a tremendous success. We are thankful for the opportunity.
The source is available at XCFD github
The team worked on a code developed by Adrian Harwood which solves problems in Computational Fluid Dynamics (CFD) using the Lattice Boltzmann Method (LBM) on multiple GPUs. A particular focus of the research group is the interaction of fluids with embedded structures. Applications include real-time analysis of the interaction between flexible tissue and blood, energy harvesting using flexible filaments in urban areas, and urban wind analysis, e.g. natural ventilation, pedestrian wind comfort or contaminant dispersion.
Before the hackathon, the code could efficiently simulate fluids on multiple GPUs. During the hackathon, the team coded a GPU implementation of the Immersed Boundary Method (IBM) for coupling the fluid simulation to embedded structures based on a CPU implementation by Joseph O’Connor. This new functionality was then coupled to an existing structural solver developed by Benjamin Owen, which was itself made faster, and the fluid solver was coupled to an external interface developed by Marta Camps Santasmasas for coupling the LBM simulation to other simulations solving the Navier-Stokes equations. While the improvements to the code are still very preliminary, the focused nature of the hackathon format enabled rapid progress and tight interactions within the team, and everyone came away very satisfied with the progress that had been made in such a short space of time. A publication is in preparation to disseminate the new code as open source software representing a significant impact for the research group on the wider community. For more info, please read their intersteing blog post at https://researchitnews.org/2019/09/03/hacking-fluids/
The “Raining Champions” worked on the Met Office Unified Model (UM), focussing on the dynamical core. We had a cross-institutional team from NCAS-CMS, Bristol University and the Met Office.
The UM is around 30 years old and has simple data structures. The combination of descriptive directives (OpenACC kernels) and managed memory meant that the required directives were surprisingly simple and required little in the way of code modification. The hackathon was the start of a process for us. With the UM’s flat wallclock profile, good overall performance requires many routines to be ported. Going forward, we intend to further the work on the dynamical core. “Physics” sections of code – which modify the dynamical evolution of the model based on physical processes such as solar radiation – work on independent vertical columns. We intend to examine these soon, since they have a different computational characteristic to the dynamical core. We further intend to examine separately our communications layers of code, with a view to mitigating the cost of halo exchanges and the data movement they incur.
On a per-routine basis, speedups observed are – in the main – up to 10x faster compared with a full Power9 socket (20 cores); other routines are typically up to 15x slower on the GPU. These results are based on a lightweight benchmark. It’s early days!
Computational fluid dynamics (CFD) is routinely used for predicting cardiovascular-system fluid flow through medical devices such as stents. Current CFD packages ignore the suspended cellular phases of blood (red blood cells or RBCs, white blood cells and platelets) which negatively affects simulation accuracy.
A coupled CUDA-based lattice Boltzmann (LB) immersed boundary (IB) plasma-RBC solver is in development which will use a state-of-the-art spring-network-based RBC model which accounts for membrane bending resistance; the deformation and orientation of RBCs is one of the primary factors affecting blood viscosity at high shear rates or strains and simplified RBC models have led to inaccuracies in membrane properties. A discretised RBC model is shown below; the nodes joining the triangle elements can be considered to be the IB markers which interact with the fluid solver through the exchange of forces.
![Healthy blood sample constituent diagram showing ~45% RBCs (top left), illustration of stent use (top right), and representation of RBC and equivalent discretised model (bottom) [1][2]](/assets/images/gpuhack19/culbibbs-1.png)
Leveraging the CUDA framework allows for the incorporation of new state-of-the-art CUDA technologies and techniques into the code, allowing for increased performance as GPU technology matures; increased performance is a necessity for modelling RBCs in flow, due to the complexity of the cell-cell, cell-wall, and cell-plasma interactions. The test case used during the Hackathon (2D laminar flow over a quasi-rigid cylinder) is not complex but allowed for work on optimisations that lay the foundation for 3D laminar flow over a quasi-rigid sphere currently in development. A 3D test case involving a moving sphere is planned before initial coupling of the RBC model with the fluid solver.
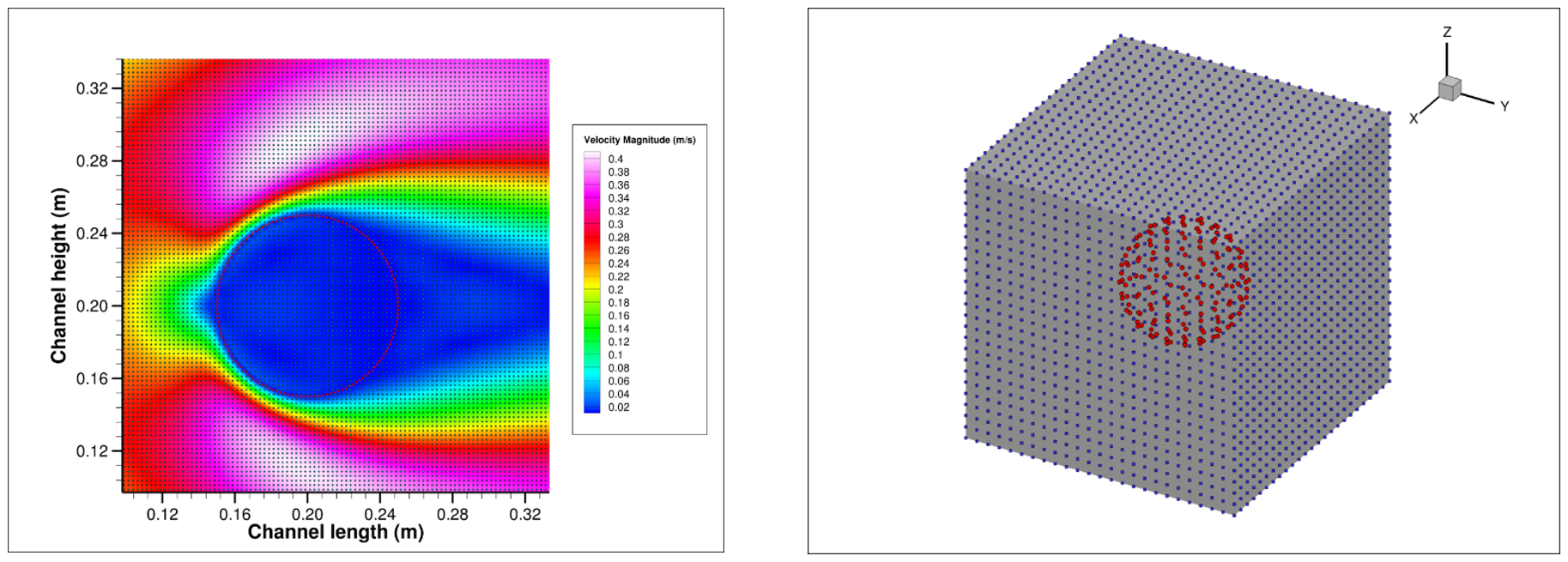
The CULBIBBS (CUDA Lattice Boltzmann Immersed Boundary Blood Solver) team comprised of TU Dublin PhD Researcher Gerald Gallagher, Irish Centre for High-End Computing (ICHEC) Computational Scientists Aaron Dees, Myles Doyle, Niall Moran and Oisin Robinson, and mentors NVIDIA Resident Scientist Christian Hundt and University of Sheffield Senior Lecturer and Research Software Engineer Paul Richmond.
The expertise in the team allowed for swift progression throughout the Hackathon; after initial profiling, memory layout transformations were implemented for better performance through using a struct of arrays rather than an array of structs for the IB marker set. Unified memory was also introduced to remove explicit data copies from host to device during the simulations and will allow for simpler implementation of multi-GPU models in the future. Following this, the serial IB functions were converted to kernels and the functionality of the force spreading from IB markers to LB nodes was modified to be IB-marker-centred rather than LB-node-centred. Atomic addition operations were used as multiple threads working on multiple IB markers need to concurrently write interpolated force data to the same LB node creating possible race conditions.
Overall, excellent speedups of up to 200x were achieved, and the following plot showing near-linear scalability was obtained.
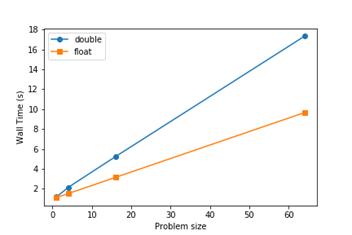
The collaborative environment at the Hackathon was fantastic and led to discussions with the team from the University of Manchester after the Hackathon. The work following this involved implementing an upgraded multi direct-forcing IB method including the optimisations from the Hackathon instead of the simpler penalty force IB method, and analysis of the atomic addition operations to completely validate the fully parallelised model. Discussions about the use of GPU interconnect technologies for future multi-GPU models and sample NVLink implementations contributed during the Hackathon will massively aid future work.
HashMaps and other related concepts were also discussed as efficient methods of storing and accessing information about the LB nodes involved in interactions with IB markers. Using this, the force spreading might be able to be changed back to LB-node-centred and the atomic addition operations removed. This has a natural application to stationary IB markers but could also have an application as the number of deformable IB RBC models increases and move through the flow domain. A necessary initial search through all LB nodes for LB IB interactions to build the HashMap could be reduced to a smaller search at each time step through knowledge about the forces, velocities and positions at the IB markers and updates to the HashMap. This would lead to further performance increases.
[1] Blausen.com, “Medical gallery of Blausen Medical 2014”. WikiJournal of Medicine 1 (2). DOI:10.15347/wjm/2014.010. ISSN 2002-4436, 2014
[2] M. Chen and F. Boyle, “Investigation of membrane mechanics using spring networks: Application to red-blood-cell modelling”, Materials Science and Engineering: C, vol. 43, pp. 506-516, 2014
The source is available at culbibbs github.
With a total of 45 people in attendance, the event was very well run and received and all this progress would not have been possible without the mentors that supported the different teams. Many thanks go to them as well as to their home organizations: NVIDIA, CSCS Swiss National Supercomputing Centre, Helmholtz-Zentrum Dresden-Rossendorf (HZDR), Science and Technology Facilities Council (STFC), Irish Centre for High-End Computing (ICHEC), Oak Ridge Leadership Computing Facility (OLCF), University of Oxford, Edinburgh Parallel Computing Centre (EPCC), and the University of Sheffield.
For further information and to apply, visit http://gpuhackathons.org/.
For queries relating to collaborating with the RSE team on projects: rse@sheffield.ac.uk
Information and access to Bede.
Join our mailing list so as to be notified when we advertise talks and workshops by subscribing to this Google Group.
Queries regarding free research computing support/guidance should be raised via our Code clinic or directed to the University IT helpdesk.
List of archived pages: Archive.