 Code written for research, and particularly in an academic context can (by my own admission…) often be a mess of comments, temporary sections, and poorly-named variables that only runs correctly whilst the researcher who made it is still around. This, despite the fact that most code-writing researchers know exactly the sins we are commiting at the time. But reproducibility is one of the central tenets of science - so why in our coding have researchers been prepared to cobble together barely working assemblages of script that can often lead to unmaintainable projects?
Code written for research, and particularly in an academic context can (by my own admission…) often be a mess of comments, temporary sections, and poorly-named variables that only runs correctly whilst the researcher who made it is still around. This, despite the fact that most code-writing researchers know exactly the sins we are commiting at the time. But reproducibility is one of the central tenets of science - so why in our coding have researchers been prepared to cobble together barely working assemblages of script that can often lead to unmaintainable projects?
In modern research (do the 70s count as modern?) computing is an essential tool - perhaps even more so than the humble lab book - so how can we improve our situation and, moreover, what’s in it for us? Code is written in research for a huge variety of tasks, from instrument control to simulations to simple plotting scripts; where does best practice really apply?
In this blog post, I’ll plant the seed of what this approach really entails and what effect it can have on research, inspiring readers to dive more deeply into the topics discussed and forming a foundation for future blogs here.
The building blocks of good software practice in research.
Firstly, it’s worth introducing that when we say Research Software we aren’t really specifically refering to large commercial software projects. Really what we mean is any code written in research. Research Software can be as small as a script you write to process and analyse some data for a paper, or as large as a software package used by millions of users around the world. As far as research goes, software should be written with best practice in mind regardless of its scope.
Software reproducibility, like scientific reproducibility, is the idea that computational experiments and analyses should be repeatable, verifiable and extendable by researchers other than the originators. In reality, because journal papers are the ultimate deliverable in academia, other elements often become neglected; code is jumbled together until it spits out some results and disseminating the complete code and methods often becomes an afterthought. In the ideal case, software should be: open source; portable (can be built on and runs on a variety of systems); and capable of reproducing the original results. All too often in practice the code runs once, on one researcher’s laptop, only at the point they were writing a particular table or paragraph. Reproducibility of analysis and data goes beyond being able to directly replicate results with the same software. As per the below image, The Turing Way define the replicability of analyses and data in four different ways.
The Software Sustainability Institute has a useful article on reproducible software.
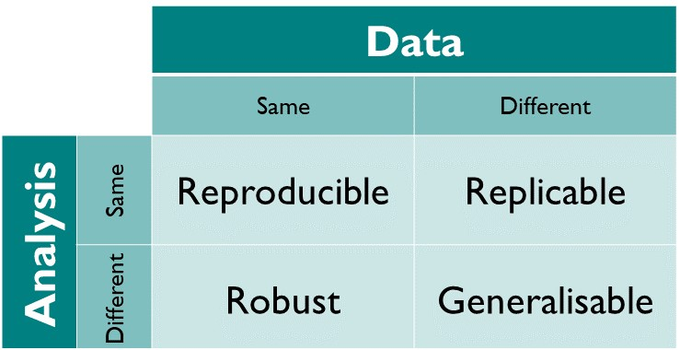
How the Turing Way defines reproducible research. (Reproduced under CC-BY from The Turing Way)
It’s all too easy to get to the point in writing software at which something runs, so we assume everything is working. By using incremental testing practices, writing programmatic tests of aspects of our software, we aim to flag these sorts of problems before they become an issue down the line. As we proceed in the development cycle, we can add additional testing on top which will examine whether any new functionality breaks anything. Software testing is a huge topic, but the essential concept is an easy idea to sell.
The Software Sustainability Institute’s’ article on testing
If you write code, then there are so many reasons to learn version control and to make your work open source. Having only recently learned to use git and GitHub for version control myself, it has been a total game-changer in the way that I write code, allowing me to write more collaboratively and accountably. Version control allows you to safely add functionality to your code, whilst protecting the main development branch. Note: git can and should be used locally for version control, not just for sharing code to GitHub or for collaboration. When developing code just for yourself on your local machine, you can use branching as a mechanism for developing new features.
Ultimately, when you release your code, being open source (maybe using a platform like GitLab or GitHub) will allow people to use your code more easily and help to solve problems, extending the reach of your coding project and facilitating future collaboration. Open source software is that for which the original source code is made freely available and may be redistributed and modified. As such, this aligns well with what we expect much of research to be: open to be learned from, developed upon, reproduced, tested and so on.
The Turing Way page on version control
Perhaps the easiest change to make to the way that you write code is just to make it better code.
A, spam and thing() are doing and can immediately understand variables and functions like speciesCount and downloadData() in your code.Examples of thrilling style guides:
Python style guide (PEP 8)R style guideMATLAB style guide
If in doubt Google probably has a style guide for your language and they are reasonable places to start.Good software isn’t just worth it for its own sake, it can have a greater influence on your research and the wider community and make your work more reproducible.
Those working on the code in future (students, collaborators, yourself) will have a much easier and more-productive time, ultimately getting more done and resulting in better research. By making your code easier to read and develop, your future students, postdocs and self won’t have to waste time rummaging through a mess of code or reinventing the wheel. With a version control system, future workers can fork the original codebase, develop new functionality, before simply merging back into the main branch of the code. Platforms such as GitLab and GitHub also facilitate accountable communication and review procedures as well as project management tools, giving you oversight over your research code and recording the changes made.
Rather than keeping your code secret and only intelligible to yourself, why not make it available online and understandable to others? That way, someone can repeat your analysis and spawn a new research topic using the methods you have developed, citing your work in the process and making it a more important element of the field. Doesn’t that sound great?! By using software best practice your code will live on beyond the time you initially have to spend on it, influencing your field and other researchers and ultimately leading to more impactful research - a valuable comodity in today’s research environment. If you need to keep your work under wraps before publication, it’s possible to just publish the code to an online repository once you’re ready. Version control systems can be run on your local machine if it’s just you working on your code, or perhaps on an institutional server for you and your collaborators to use as your remote repo during development before publishing.
So you’ve bought into the idea that software best practice is a productive use of your team’s time, but how do you actually implement it in reality? Is it truly possible to practically do this in a research context? Of course, the answer is yes! and plenty of researchers are doing it. But you aren’t alone and there are a fantastic number of resources to help you to do this.

Learning version control using git or another system is one of the best changes you can make to the way you code. Though it can be daunting at first (I’ve only recently learned myself), ease comes with practice. There are loads of ways to learn:
Further, version control through git can be done in a graphical IDE like Rstudio, MATLAB, Visual Studio Code et al. or GitHub Desktop or even some text editors such as atom.
In platforms such as GitLab and GitHub, open source development is very easily handled, simply by making the code available on a public repository, anyone can contribute to your project, with your control easily maintained via setting permissions. Contributors are encouraged to ‘fork’ code (make their own copy) before making changes, after which you can choose whether to merge their changes into your master codebase. Training is provided online via GitHub or take part in a Software Carpentry-style workshop to learn these skills and others.
How to improve the reproducibility of your research is a huge question, but definitely surmountable. It is certainly worth ensuring that your code can be reproduced outside of your lab so that it can have a lifetime and influence beyond one research paper. What writing a reproducible piece of code really entails depends on the scale of your project, it could be as simple as putting your analysis into an R notebook to make available online alongside a paper (or other method of disemination); or for a larger piece of code then containerisation could be a useful tool (recent RSE blog article on containers). Some more reproducible code resources:
Perhaps this all sounds like something that you need a dedicated specialist for, or maybe just somewhere that you need advice? This is where RSEs come in. Depending on the level of assistance you need, it’s possible to either:
Of course, there is some personal bias here, but RSEs really are tasked with helping to deliver improved research software in a variety of ways, with the express aims of improved reproducibility, increased software impact and code with a more influential lifetime.
git with GitHub Dr David Wilby is the newest recruit to the RSE team. Coming from a background in physics and sensory biology, he has previous experience in using numerical simulation to interrogate animal visual anatomy. Other coding experience includes writing instrument control software and data analysis. His other non-coding research experience includes training chickens and scaring hermit crabs.
In the RSE team, David’s main role is to undertake work in developing the impact of software from the Department of Computer Science at the University of Sheffield.
Dr David Wilby is the newest recruit to the RSE team. Coming from a background in physics and sensory biology, he has previous experience in using numerical simulation to interrogate animal visual anatomy. Other coding experience includes writing instrument control software and data analysis. His other non-coding research experience includes training chickens and scaring hermit crabs.
In the RSE team, David’s main role is to undertake work in developing the impact of software from the Department of Computer Science at the University of Sheffield.
For queries relating to collaborating with the RSE team on projects: rse@sheffield.ac.uk
Information and access to Bede.
Join our mailing list so as to be notified when we advertise talks and workshops by subscribing to this Google Group.
Queries regarding free research computing support/guidance should be raised via our Code clinic or directed to the University IT helpdesk.
List of archived pages: Archive.